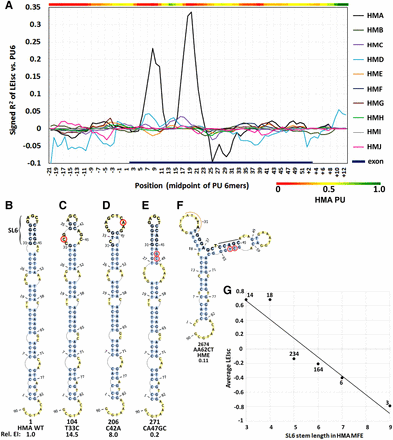
A stem–loop secondary structure in the HMA sequence inhibits splicing. (A) Map of the correlation (signed R2) of splicing with the probability of being unpaired (PU) in secondary structures. For each starting position of each HM, each mutant window of 6 nt was evaluated for its average PU. These PU values were then correlated to LEIsc scores. Note that the true WT HMA exhibits a strong region-specific correlation from exonic positions 8 to 22 but the other nine HMs show no such strong effect at any position. The heat map shows the PU values of WT HMA 6-mers. (B–F). The minimum free energy (MFE) structures of selected 90-nt folded sequences. Mutant serial numbers are indicated for reference to Supplemental Table S2.The 15-nt sequence of HMA stem–loop 6 is in bold. The base changes and the EI value relative to the WT are indicated. (B) Wild-type HMA. The location of stem–loop 6 (SL6), the structure that inhibits splicing, is indicated (bases 31 to 45); its coordinates in the exon are 8 to 22. (C) The MFE structure of an HMA mutation (circled) with a SBS in the upstream arm of SL6. (D) As in C, but the mutation is in the downstream arm of SL6. (E) An example of an HMA DBS that extends the stem length of SL6 from 5 to 9. This more stable stem produces a further reduction in splicing (panel G). (F) A contrasting example of a different HM, HME. The location of the hexamer difference from exon positions 5 to 10 is shown by the orange arc. The black bar indicates the sequence of what is the downstream arm of SL6 in HMA. Note the largely different MFE structure compared to HMA. (G) The double-strandedness of the SL6 stem of HMA correlates with splicing. The points show the average LEIsc values for MFE structures having the indicated number of paired bases in the SL6 stem. The points are labeled with the number of mutants having that stem length.











