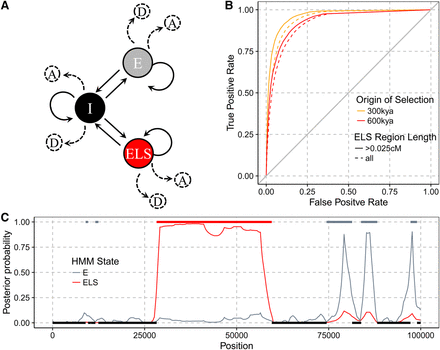
(A) Graphical representation of the Extended Lineage Sorting hidden Markov model. States are depicted by nodes and transitions by edges. Each state emits an archaic allele as either derived, D, or ancestral, A, depending on the type of site in the modern human population (fixed or segregating at a given frequency). States are labeled I for Internal, E for External, and ELS for Extended Lineage Sorting. (B) Receiver operator curves for varying cutoffs on the posterior probability of the ELS state and counting the number of sites in ELS regions that were correctly labeled. All bases labeled ELS outside of simulated ELS regions are considered false positives. Sites in ELS regions with a posterior probability below the cutoff are considered false negatives. (C) Example of the labeling of a simulated ELS region. Horizontal bars indicate true external (top) and internal (bottom) regions. The posterior probability is shown in red for ELS regions and in gray for E regions. The region overlapping position 50,000 (red bar) is caused by a simulated selective sweep.











