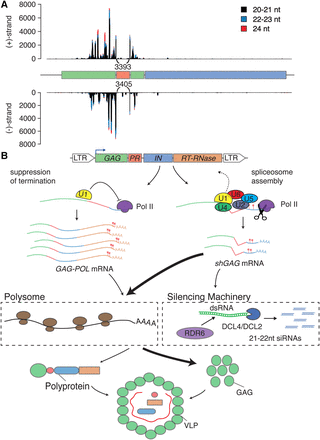
Spliced subgenomic shGAG mRNAs display many termination codons in long 3′ UTRs, and are specifically targeted by the RNA silencing machinery. (A) We mapped 20- to 24-nt siRNA profile from the EVD locus in epi15 F11 plants mapped with a splice-aware mapping software. Arcs display previously unmapped splice junction siRNA reads. (B) A model showing two concurrent modes of action of the U1 snRNP either repressing premature termination to allow full-length mRNA/genomic RNA synthesis or promoting splicing and subsequent premature termination of the shGAG mRNA. The two generated isoforms, shGAG and GAG-POL, differentially associate with polysomes, allowing proper regulation of protein abundance required for successful VLP formation and transposition. The shGAG mRNA also specifically activates the host silencing response, depicted here in its primary PTGS phase involving 21- to 22-nt siRNAs produced by DLC4 and DCL2, respectively. Note that protection of EVD RNAs due to GAG particle formation allows them to resist PTGS at this stage, as shown by Marí-Ordóñez et al. (2013).











