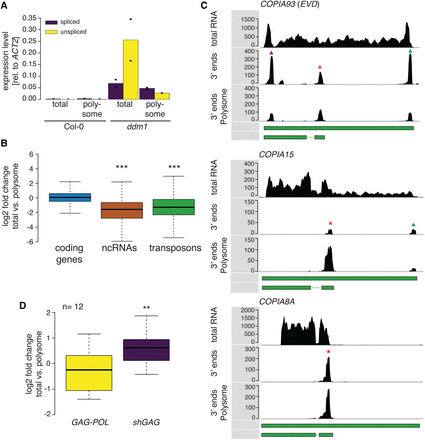
The two Ty1/Copia mRNA isoforms differentially associate with polysomes. (A) Relative steady-state levels of EVD spliced and unspliced isoforms found in the total as opposed to polysome fractions. Dots indicate measurements made in the two individual replicates. (B) Log2 accumulation fold change between total RNA and polysome-associated RNA of various RNA classes. (C) RNA 3′ end sequence coverage of mRNAs from total cell extract and polysome-associated mRNAs, showing differential loading of shGAG (red asterisk) and GAG-POL (green triangle) mRNAs of EVD and COPIA15. Note that an additional peak is visible in the EVD 5′ LTR owing to high sequence homology between the two LTRs (purple triangle). (D) Differential association of Ty1/Copia shGAG mRNA and GAG-POL mRNA to polysomes. Read coverages 50 nt downstream from and 300 nt upstream of the respective termination sites were taken into account for quantification. (**) P < 0.01, (***) P < 0.001 (Wilcoxon rank-sum test); (n) number of individual elements analyzed.











