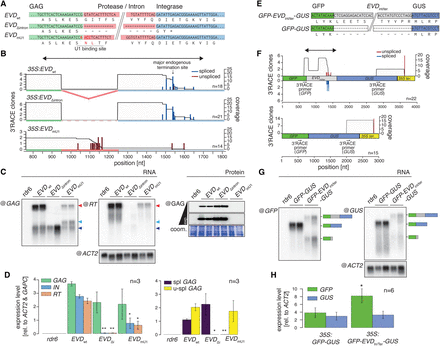
Mutational analyses of ectopically expressed EVD and full reconstruction of its splicing behavior in a reporter system. (A) Nucleotide and corresponding amino acid sequence of 35S:EVDwt, 35S:EVDΔintron, and 35S:EVDmU1. (B) Individual 3′ RACE clones from distinct 35S:EVDwt, 35S:EVDΔintron, and 35S:EVDmU1 overexpression lines in the rdr6 background. Blue and red bars display spliced and unspliced 3′ ends and gray areas the cumulative sequence coverage, similar to Figure 3C. Positions are indicated in nucleotides (nt) from the EVD start codon. (C) RNA and protein blot analysis of 35S:EVDwt, 35S:EVDΔintron, and 35S:EVDmU1 overexpression lines in rdr6. Total RNA was consecutively hybridized with RT- and GAG-specific probes (see Fig. 3A) to identify the GAG-POL RNA (red arrow) and shGAG RNA (blue arrows), respectively. The spliced shGAG RNA and an unspliced RNA cryptically terminated inside the intron comigrate (dark blue arrow); a size shift is visible for the unspliced RNA terminating at the endogenous GAG terminator (light blue arrow). (Bottom panel) Western blot analysis conducted with the anti-GAG antibody. (ACT2) ACTIN2 mRNA loading control; (coom.) Coomassie staining of total protein as a loading control. (D) qRT-PCR measurements of the relative expression levels of 35S:EVDwt, 35S:EVDΔintron, and 35S:EVDmU1. (E) Nucleotide and corresponding amino-acid sequence of 35S:GFP-EVDIn/Ter-GUS and 35S:GFP-GUS. (F) 3′ RACE analysis of 35S:GFP-EVDIn/Ter-GUS and 35S:GFP-GUS in the rdr6 background showing spliced and unspliced 3′ ends as in B. The double-headed arrow indicates the EVDIn/Ter sequence also depicted in B. Positions are indicated in nucleotides (nt) from the GFP start codon. (G) Northern blot analysis of alternative RNA isoforms produced from both reporters. The blot was consecutively hybridized with GFP- and GUS-specific probes to detect GFP and full-length GFP-GUS RNA species, respectively. ACTIN 2 (ACT2) mRNA loading control. (H) Relative expression levels of GFP and GUS RNA indicate modulation of GFP-to-GUS ratios. In all graphs, (*) P < 0.05; (**) P < 0.01 (two-sided t-test against corresponding controls); (n) number of biological replicates or individual 3′ RACE clones sequenced; error bars, SE.











