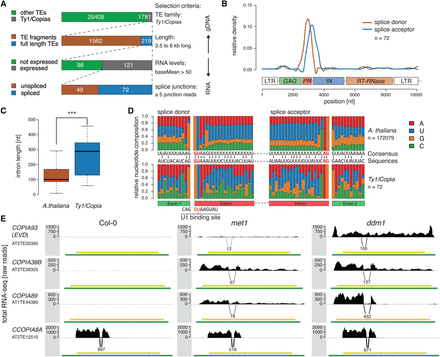
Genome-wide analysis of splicing of Ty1/Copia elements in A. thaliana. (A) Selection of spliced Ty1/Copia elements from RNA-seq data of globally reactivated transposons. (B) Density profile of splice donor and splice acceptor sites on a multiple sequence alignment of spliced Ty1/Copia elements selected in A. (C,D) Intron length (C) and (D) nucleotide sequence composition (D) of detected Ty1/Copia introns and A. thaliana gene introns. (E) RNA-seq profiles of spliced Ty1/Copia elements in wild-type (Col-0), met1, and ddm1 backgrounds. A continuum of splicing intensities is depicted with specific examples, displayed by arcs and numbers below the sequence coverage indicating splice-junction reads. Green annotation bars represent the entire TE annotation, including LTRs, whereas yellow annotation bars indicate the TE gene annotation covering only the protein coding sequence. In all panels, (***) P < 0.001 (Wilcoxon rank-sum test); (n) number of individual elements or introns analyzed.











