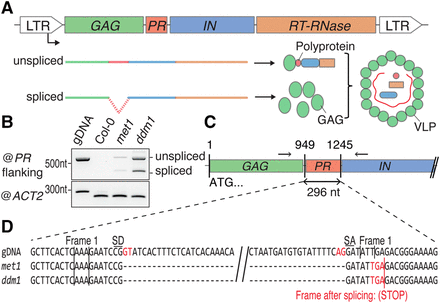
A scheme for Ty1/Copia protein production and alternative splicing of the EVD RNA. (A) Scheme of the generic Ty1/Copia elements’ genome and a putative alternative splicing strategy to modulate protein abundances from the condensed genome, as required for successful TE life cycle. (B) Alternative isoforms of EVD detected by RT-PCR using primers flanking the protease domain. ACTIN 2 (ACT2) PCR uses primers flanking an intron to amplify a longer PCR product corresponding to the genomic DNA (gDNA) and a shorter cDNA form. It serves as a loading control and validates the absence of genomic DNA contamination. (C) Schematic representation of the region surrounding the EVD intron. Arrows indicate the flanking primers used in B. (D) Genomic (gDNA) and spliced sequences of the region flanking the EVD intron in met1 and ddm1 backgrounds. (LTR) long terminal repeat; (PR) protease; (IN) integrase; (RT-RNase) reverse-transcriptase-RNase; (VLP) viral-like particle; (SD) splice donor; (SA) splice acceptor.











