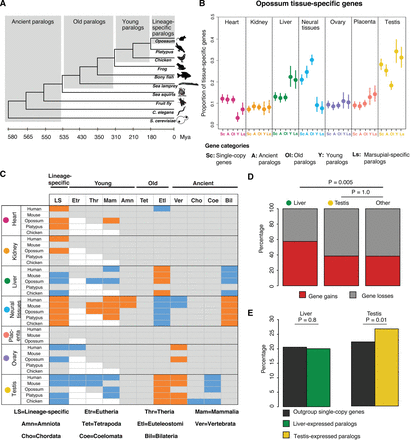
Evolutionary dynamic expression profiles of tissue-specific expressed genes. (A) Schematic representation of duplication age categories using opossum as an example. (B) Proportion of genes specific to any given tissue is plotted by gene type and duplication age. Tissue specificity of opossum genes was assessed using the “stringent” definition (see Methods). Single-copy genes and paralogs, grouped in four age classes, are shown for each tissue. Bars represent 95% confidence intervals. Analyses carried out at gene level. (C) Significant differences in evolutionary dynamics of tissue preferences of duplicated genes. Duplication ages are given in the upper row. Bars above group the fine-grained duplication ages into the same four age categories as in A (Methods). Significant over- (blue) or underrepresentation (orange) of genes with highest expression in a given tissue was tested for paralogs in each species and for each duplication age with a χ2 test, followed by a post hoc procedure (Methods). Gray cells signify no statistical difference. (D) Analysis of differential gene loss/gain in gene families with highest expression in liver, testis, and all other tissues combined. (E) Expression shift analysis: Proportions of chicken outgroup genes with highest expression in liver and testis compared to the proportion of resulting mammalian paralogs with highest expression in these tissues. Significance was assessed with a χ2 test.











