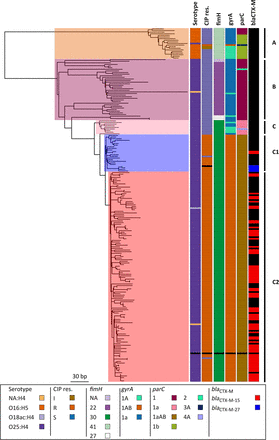
ST131 maximum-likelihood phylogenetic tree based on SNPs called against the reference EC958. Columns to the right of the tree show the in silico predicted serotype (O16-H5 or O25-H4); phenotypic resistance to ciprofloxacin (CIP res.); SNP-based definition of fimH, gyrA, and parC genotypes; and the presence of blaCTX-M and the type. NA:H4 in the serotype indicates that we were unable to assign a definite O type for the isolate. It has not been counted as a new serotype. Clades assigned based on the markers and clade-specific SNPs are shown on the right. The only isolate with missing data (black) is the reference strain EC958. The tree is mid-point rooted.











