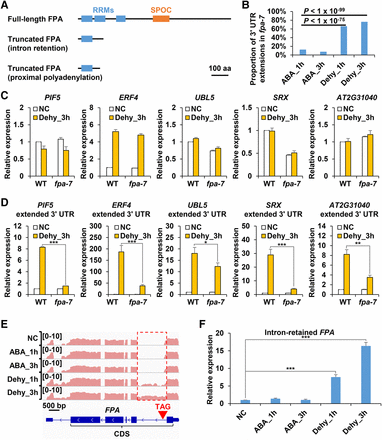
FPA partially regulates biogenesis of 3′ UTR extensions. (A) Schematic diagram showing domains of the full-length FPA protein (top), intron retention-produced truncated FPA protein (middle), and proximal polyadenylation-produced FPA protein (bottom). (B) Proportion of ABA treatment- and dehydration stress–induced 3′ UTR extensions among 3′ UTR extensions induced in fpa-7 mutant. (C) Real-time PCR analysis of total expression levels of PIF5, ERF4, UBL5, SRX, and AT2G31040 in WT and fpa-7 mutant. The expression level of each gene in WT plants grown in normal condition (NC) was set to 1. (D) Real-time PCR analysis showing induction of 3′ UTR extensions by dehydration stress of PIF5, ERF4, UBL5, SRX, and AT2G31040 in WT and fpa-7 mutant. The expression levels of each gene in WT plants and fpa-7 mutants grown in normal condition (NC) were set to 1. (E) The IGV Genome Browser view showing FPA. Pink color represents read abundance of reverse strand. Retained intron is depicted by a red rectangle. The position of introduced premature termination codon is indicated by a red inverted triangle. (F) Real-time PCR analysis of expression levels of intron-retained FPA. The expression level in normal condition (NC) was set to 1. Transcript levels were normalized to ACT2 expression. Data are shown as mean ± SD (n = 3) of one representative result of three independent biological replicates. Asterisks indicate statistically significant differences compared with WT: (*) P < 0.05; (**) P < 0.01; (***) P < 0.001; t-test.











