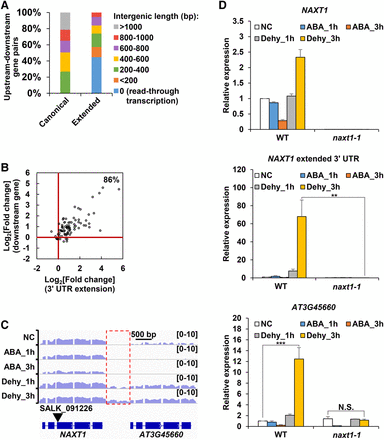
Activation or read-through transcription of downstream genes by 3′ UTR extensions. (A) Composition of upstream–downstream gene pairs of canonical 3′ UTRs and extended 3′ UTRs. A total of 429 gene pairs were used for the plot. (B) Fold change of expression levels (FPKM) of 3′ UTR extensions (x-axis) versus fold change of expression levels of sense genes (y-axis) in Dehy_3h compared with Dehy_1h. Log2-transformed fold changes were used for this plot. (C) The IGV Genome Browser view showing NAXT1 and AT3G45660. Blue color represents read abundance of forward strand. Extended 3′ UTRs is depicted by a red rectangle. T-DNA insertion site is indicated by a black inverted triangle. (D) Real-time PCR analysis of expression levels of NAXT1 (top), NAXT1 extended 3′ UTR (middle), and AT3G45660 (bottom) in WT and naxt1-1 mutant. Transcript levels were normalized to ACT2 expression. The expression level of each gene in WT plants grown in normal condition (NC) was set to 1. Data are shown as mean ± SD (n = 3) of one representative result of three independent biological replicates. Asterisks indicate statistically significant differences: (**) P < 0.01; (***) P < 0.001; t-test; (N.S.) not significant.











