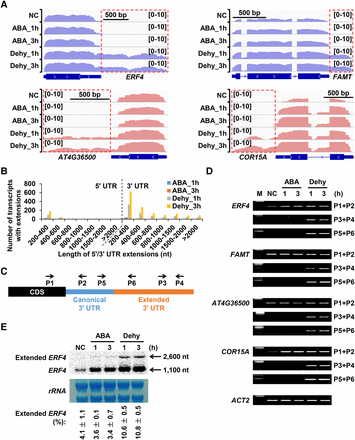
Dehydration stress induces 3′ UTR extensions. (A) Examples of dehydration stress–induced 3′ UTR extensions. Blue and pink colors represent read abundance of forward strand and reverse strand, respectively. Extended 3′ UTRs are depicted by red rectangles. (B) Number of transcripts with 5′/3′ UTR extensions in ABA treatment and dehydration stress. (C) Schematic diagram of RT-PCR primers used to validate 3′ UTR extensions. (D) Validation of dehydration stress–induced 3′ UTR extensions (A) using RT-PCR. Primer set P1+P2 indicates transcripts with canonical and extended 3′ UTRs. Primer sets P3+P4 and P5+P6 indicate extended 3′ UTRs. ACT2 mRNA served as a positive control. (E) Validation of ERF4 3′ UTR extension using RNA gel blot analysis. 32P-labeled riboprobe targeting ERF4 3′ UTR region was used to detect both the canonical and extended ERF4 transcripts. Methylene blue–stained ribosomal RNA (rRNA) is shown as loading control. The percentage of extended ERF4 in total ERF4 transcripts are presented as mean ± SD of two biological replicates.











