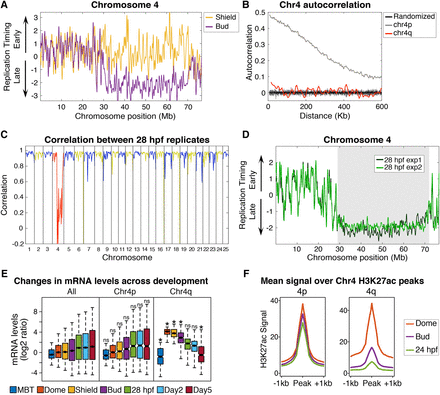
The long arm of Chromosome 4 undergoes a developmentally regulated switch to late replication. (A) Replication timing of Chromosome 4 plotted at Shield (gold) and Bud (purple) stages. (B) Autocorrelation of replication timing for a 25-Mb segment of the late replicating long arm of Chromosome 4 (Chr 4q, red), a 25-Mb segment of the short arm of Chromosome 4 (Chr 4p, gray), and randomly permutated data (black lines). (C) Correlation between biological replicates of 28 hpf embryos. Blue and yellow denote alternating chromosomes, red highlights Chromosome 4. (D) Replication timing profiles for 28 hpf embryos show a low consistency in replication timing between experimental replicates along the long arm of Chromosome 4. The gray box indicates a region of low correlation between experimental replicates (average r = 0.16). (E) mRNA levels (log2 ratio of fold change from transcript counts at the two-cell stage) for RefSeq genes on all chromosomes except Chromosome 4 (All), genes on the short arm of Chromosome 4 (Chr 4p), or genes on the late replicating long arm of Chromosome 4 (Chr 4q): (*) comparison to All genes at the matching developmental stage, two-way ANOVA with Bonferroni corrected P-value <0.005. Box plots show the median (line), 95% confidence interval (notch), 25th–75th percentile (box), and 10th–90th percentile (whiskers). (F) Average H3K27ac signal ±1 kb from all Dome peak apexes on the short (4p) and long (4q) arms of Chromosome 4 at Dome (orange), Bud (purple), and 28 hpf (green) (Pauli et al. 2012).











