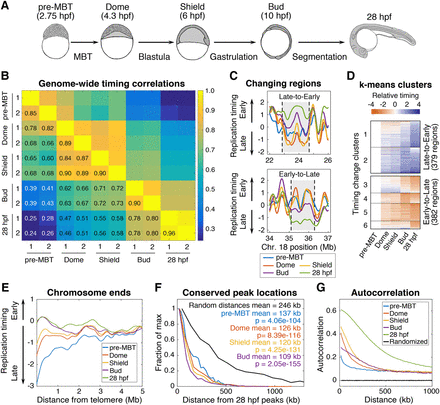
Replication timing is dynamically regulated throughout development. (A) Developmental time points used in this study. (B) Pearson's correlation matrix comparing genome-wide replication timing values from biological replicates of each developmental time point to all other samples. (C) Using a two-state hidden Markov model (HMM) the whole genome was segmented into 1620 contiguous regions that either change timing from early-to-late or late-to-early throughout development. Representative genomic regions that switch from early-to-late and late-to-early are shown. (D) Using k-means clustering, the HMM-defined changing regions were grouped based on the patterns of their timing changes. (E) Replication timing plotted as a function of distance from the telomere for each developmental time point. (F) Distances between the location of peaks in the 28 hpf sample and peaks in all other developmental samples (P-values from Kolmogorov-Smirnov test). (G) Autocorrelation of replication timing for each developmental sample (colored lines) and randomly permutated data (black lines).











