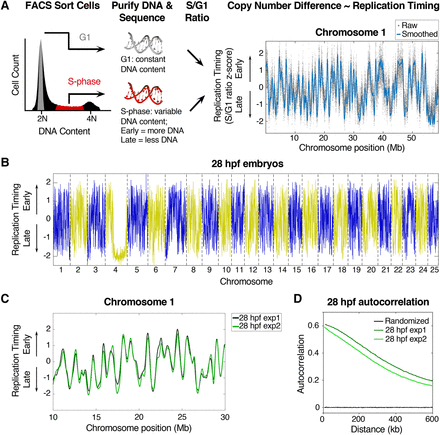
High-resolution replication timing analysis in zebrafish. (A) Experimental outline: cells were FACS sorted based on DNA content into G1 and S-phase fractions, DNA was purified and sequenced, and replication timing was calculated based on variations in DNA copy number between G1 and S-phase cells (S/G1 ratio). To normalize, the genome mean replication timing value was set to 0, and the standard deviation was set to 1 (for full details, see Supplemental Figure S1A; Methods and Supplemental Methods). (B) Whole-genome replication timing profile for 28 hpf zebrafish embryos: (blue/yellow) odd/even chromosomes. (C) Replication timing profiles for two experimental replicates of 28 hpf embryos. (D) Autocorrelation of replication timing values along the length of the chromosomes (except Chromosome 4) for two experimental replicates of 28 hpf embryos.











