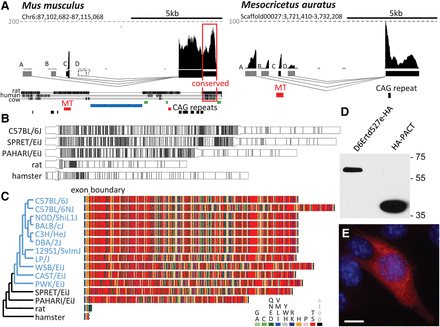
A solo MTD LTR contribution to de novo evolution of a protein-coding gene. (A) Genomic organization of the D6Ertd527e locus in Mus musculus and Mesocricetus auratus. Shown are UCSC Genome Browser snapshots of D6Ertd527e loci with mapped oocyte RNA NGS. The gray dashed lines indicate CPMs. Below the conservation track is the RepeatMasker track with MT LTR insertions in red, SINE insertions in green, and a large LINE-1 insert in blue. The conserved 3′ UTR region is framed. (B) CAG trinucleotide density in MTD-driven transcripts in mice, rat, and hamster. Each CAG is represented by a vertical line. The widening depicts the coding sequence; the initiation codon is in the MTD exon. (C) Virtual translation of MTD-driven D6Ertd527e transcripts from rodent species (black) and mouse strains (blue). The phylogenetic tree was adopted from Nellaker et al. (2012). (D) D6ERTD527E protein expression in NIH3T3 cells. Transiently transfected cells expressing C-terminally HA-tagged D6ERTD527E or N-terminally HA-tagged PACT (control) were analyzed 48 h post-transfection by immunoblotting. (E) Ectopically expressed C-terminally HA-tagged D6ERTD527E protein (red) has cytoplasmic localization in mouse NIH3T3 cells. DNA (blue) was stained with DAPI. Untransfected cells lacking the HA signal demonstrate staining specificity. Scale bar, 10 μm.











