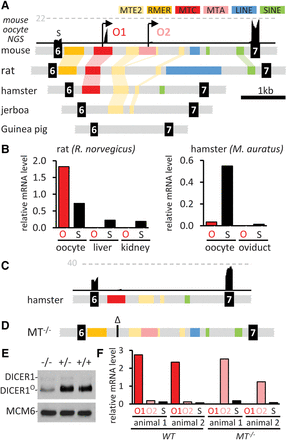
Dicer1 rewiring and remodeling by MT LTRs. (A) Retrotransposon content changes during evolution of Dicer1 intron 6 in rodents. Above the mouse sequence is a snapshot of a UCSC Genome Browser track with mouse oocyte NGS data. The gray dashed line indicates CPM. O1, O2—two oocyte-specific promoters. (B) qPCR analysis of Dicer1 isoform mRNA expression in rat and hamster oocytes. Dicer1O (O) and full-length somatic Dicer1 isoform (S) expression are shown relative to Hprt. (C) NGS data support minimal Dicer1O expression in hamster oocytes. Shown is a UCSC Genome Browser snapshot. The horizontal dashed line represents the number of reads. (D) A schematic view of the intron 6 in Dicer1MT−/− mice with MTC (O1 promoter) deletion. (E) Oocytes lacking the MTC LTR (O1 promoter) still produce a detectable amount of DICER1O. Shown is an immunoblot from C57Bl/6NCrl oocytes. A low amount of the full-length DICER1 is visible above the DICER1O isoform. Each lane represents roughly 500 oocytes. (F) qPCR analysis of Dicer1 transcripts driven by MTC (O1) and MTA (O2) LTRs. Dicer1 expression is shown relative to Hprt.











