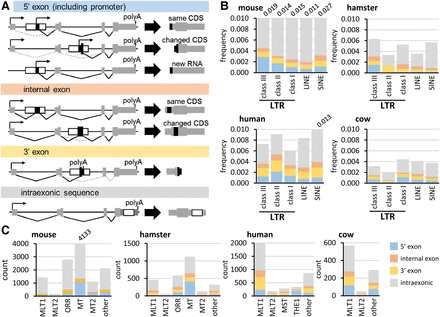
Gene remodeling by LTRs. (A) Four categories of LTR co-option according to the co-opted exon boundaries. LTR co-options may affect gene expression but not the encoded protein, remodel a gene and change its protein product, or create a new transcriptional unit, such as an lncRNA gene. (B) Whole-genome analyses of impacts of LTR, LINE, and SINE elements on gene structure according to the classification depicted in A. Repeatmasker (Smit et al. 2013–2015) was used for Class I–III LTR annotation. The y-scale depicts the ratio of observed co-option events and annotated insertions, which are listed in Supplemental Tables S2 (mouse), S5 (hamster), S6 (human), and S7 (cow). (C) Impact of MaLR and MT2 LTRs on gene structure according to the classification depicted in A in four mammals. The y-scale depicts the number of co-opted insertion events. B and C display both full and partial contributions.











