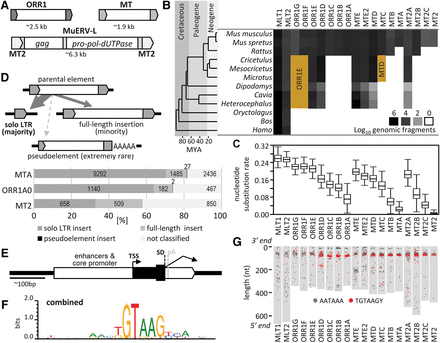
Sequence properties of selected ERVL LTRs. (A) Organization of ORR1, MT, and MuERV-L retrotransposons. Internal sequences of ORR1 and MT elements do not encode any protein. (B) Abundance of selected ERVL LTRs in mammalian genomes. The brown areas indicate misannotated ORR1F, ORR1G, and MTC LTRs in genomes of other rodents. (C) Nucleotide substitution rate for the closest pairs among 200 random inserts in each LTR subfamily. (D) Three types of LTR retrotransposon inserts and their frequencies among the selected youngest ERVL subfamilies. (E) A schematic depiction of an MT LTR gene-remodeling platform. (F) A combined SD sequence logo of MT, ORR1, and MT2 LTR families. (G) Conserved position of the splice consensus sequence at the 3′ end of selected LTRs. Gray rectangles depict consensus lengths of LTRs aligned by the 3′ end to the top. Red or black points represent positions of TGTAAGY consensus motif or AATAAA polyadenylation signal, respectively, in 200 randomly chosen LTRs in each subfamily.











