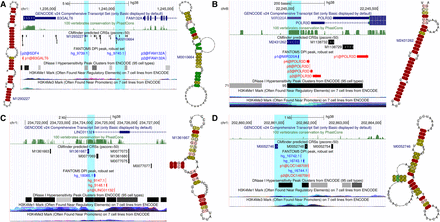
Example CRSs in gene regulatory regions are supported by unannotated transcript boundaries. (A) Two intergenic enhancers in a highly structured region of low SI (CRS M1293227 is only conserved in primates) between two gene 3′ ends. (B) MIR320A is upstream of POLR3D TSS. (C) Anti-sense transcription at the promoter of the lncRNA LINC01132 has enhancer-like chromatin signatures. (D) Intergenic enhancer with unidirectional stable transcription from the minus strand as measured by control and exosome-depleted HeLa cells (Andersson et al. 2014b). Color code in consensus structures is the level of base-pair conservation in the structure-based alignments.











