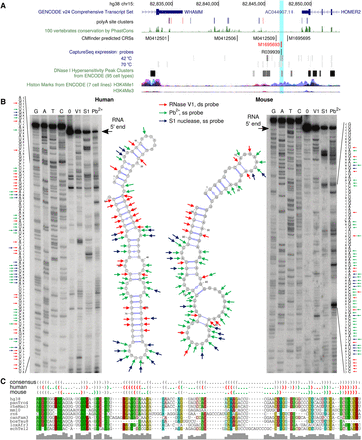
In vitro RNA structure probing in human and mouse shows conserved structure of CRS M1695693. FDR is 11.0% and SI of the nine-species (filtered from the 17-species tree) structural alignment is 48% (45% between human and mouse). The CRS is located between the 3′ UTRs of HOMER2 (minus strand) and WHAMM (plus strand; Chr 15: 8284671–82846804). It overlaps a DNase hypersensitive site (DHS) (ENCODE) and has the typical chromatin signatures of enhancers, namely, enrichment of H3K4me1 and reduced enrichment of H3K4me3, all indicators for a transcribed regulatory region. However, CAGE data from FANTOM5 did not support this hypothesis; instead, poly(A) site clusters (Gruber et al. 2016) suggest an extended 3′ UTR of HOMER2. (A) Genomic tracks. (B) Structure probing results in human and mouse, where red marks base-paired nucleotides (ds), and green and blue mark single-stranded nucleotides (ss). (C) CMfinder's structural alignment, predicted consensus RNA secondary structure, and predicted individual structures in human and mouse as dot-bracket notation. The probing results are overlapped with the in silico predictions by their color code.











