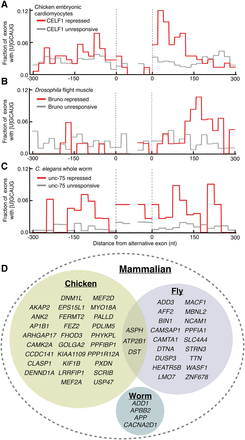
RBFOX motifs are enriched downstream of exons repressed by ancient CELF homologs. Histogram of the frequency of [U]GCAUG motifs in nonoverlapping 20-nt windows within 300 nt of alternative exons that are repressed by ancient CELF family homologs (red) or were unresponsive to depletion/mutation (gray) in (A) chicken embryonic cardiomyocytes (control versus siRNA knockdown of CELF1) (Blech-Hermoni and Ladd 2015), (B) Drosophila melanogaster indirect flight muscles (control versus RNAi hairpin knockdown of bru1) (Spletter et al. 2015), and (C) C. elegans whole worm (control versus unc-75 null [e950] mutants) (Norris et al. 2014). (D) Venn diagram showing mammalian gene names containing CELF/RBFOX coregulated splicing where the orthologous gene in chicken, fly, or worm also contains a putative CELF/RBFOX coregulated event (see Methods and Supplemental Table S7).











