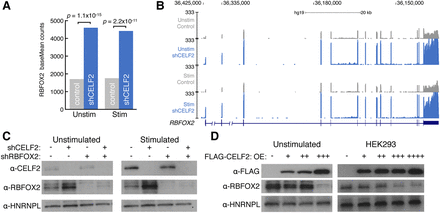
CELF2 represses RBFOX2 mRNA and protein levels in T cells. (A) RBFOX2 mRNA expression in unstimulated and stimulated JSL1 T cells (gray) compared to CELF2 depletion (blue) as determined by DESeq. Significant increases in RBFOX2 mRNA levels upon CELF2 depletion was determined by DESeq (fold-change > 2.5 and adjusted P < 2.3 × 10−11). (B) UCSC Genome Browser snapshot of RNA-seq reads over the RBFOX2 locus in unstimulated (top) or stimulated (bottom) control (gray) versus CELF2 depletion (blue) JSL1 cells. (C) Western blot monitoring protein levels of CELF2 and RBFOX2 in unstimulated (left) or stimulated (right) JSL1 cells depleted for CELF2 (lane 2), RBFOX2 (lane 3), or both (lane 4). An antibody against HNRNPL was used as a loading control. (D) Western blot monitoring FLAG-CELF2 and RBFOX2 expression in unstimulated JLS1 cells (left) or HEK293 cells (right) that expressed increasing amounts of FLAG-CELF2 construct. An antibody against HNRNPL was used as a loading control.











