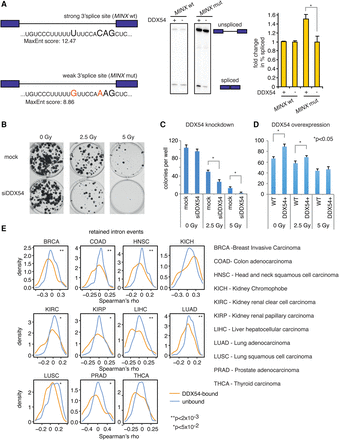
DDX54 affects in vitro splicing efficacy of weak splice site–containing transcripts, promotes cell survival, and impacts splicing of retained introns in tumor tissues. (A) A schematic of minigenes (MINX wt and mut) used for in vitro splicing assays and differing splice site strengths (left). Representative images of PAGE analysis are shown (center). Bar graph of fold changes in the percentage of PCR product corresponding to spliced versus total transcript (right). Values represent the means of two independent experiments ± standard deviations (Student's t test, * P < 0.05). (B) Representative colony formation assay of MCF-7 Flp-In cells exposed to the indicated IR doses. (C,D) Quantification of colony formation assays upon DDX54 knockdown (C) or FLAG-HA/DDX54 overexpression (D). Colonies were counted, and mean counts and standard deviations were obtained from three independent experiments (Student's t test): (*) P < 0.05. (E) Retained intron events in 11 TCGA data sets were classified as DDX54-bound or unbound according to PAR-CLIP data. Spearman correlation coefficients between expression Z-scores and PSI values in normal and tumor samples were computed, and their distributions were compared (K-S test) between DDX54-bound and unbound retained introns.











