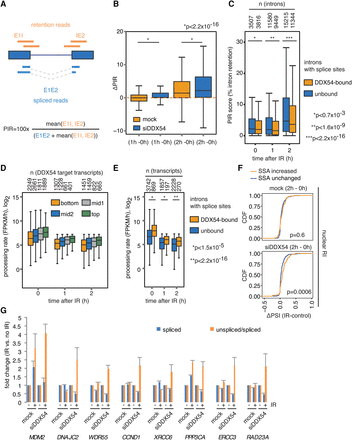
DDX54 prevents IR-induced intron retention by increasing pre-mRNA processing rates of its target transcripts. (A) Intron retention quantification by percent intron retention (PIR) scores. (B) Box plots of ΔPIR values (IR, control) obtained from 4sU-Seq data. K-S tests were used for comparison. (C) Box plot of PIR scores after different time points post IR exposure (10 Gy). Introns with at least five T-C transition events per exon–intron–exon pair were considered as DDX54-bound, and the rest as unbound. The numbers of introns per group are given above the plots. K-S tests were used for comparisons. (D) Box plot of processing rates obtained after different time periods after IR exposure (10 Gy). Transcripts were categorized according to the number of T-C transitions, and group sizes are indicated above the graph. (E) Box plots of processing rates for transcripts with at least five T-C transition events per exon–intron–exon pair (DDX54-bound) or all other transcripts (unbound). (F) Comparison between spliceostatin A (SSA) and DDX54-regulated retained intron events. Introns with increased retention (adjusted P < 0.05, ΔPSI > 0.05) or unchanged (adjusted P > 0.05, absolute ΔPSI < 0.05) upon SSA treatment in the nucleus were identified by SUPPA. Distributions of ΔPSI values (IR, control) were compared for mock and siDDX54 conditions (Wilcoxon rank-sum tests). For the set of cytoplasmic RI, see Supplemental Figure S9D. (G) RT-PCR analysis of unspliced and spliced isoforms in newly synthesized RNA extracted from mock- or siDDX54-transfected cells that were either IR-exposed (10 Gy, 2 h) or untreated. Mean fold changes in spliced abundances and unspliced/spliced ratios and standard deviations from two determinations were plotted.











