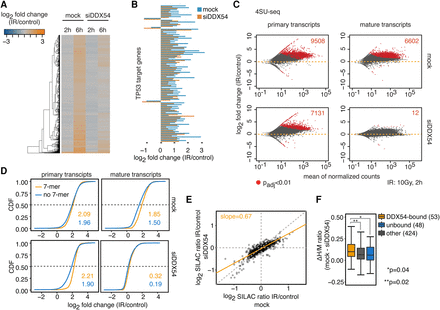
DDX54 is required for the maturation of DNA damage–induced mRNAs via a post-transcriptional mechanism. (A) Heatmap of log2-transformed fold changes (IR versus control) obtained from mRNA-seq data in mock- and siDDX54-treated cells after indicated time periods post IR exposure. (B) Bar plots of log2-transformed fold changes (IR versus control) for a selection of the most differentially expressed (absolute log2-fold changes >1 after 6 h) TP53 target genes in mRNA-seq data. The full set is shown in Supplemental Figure S8. (C) Differential primary transcript (pre-mRNA) and mature transcript (mRNA) expression analysis between IR-exposed and control MCF-7 cells that were either mock- or siDDX54-transfected. The numbers of up-regulated transcripts are indicated in the upper right corners. (D) Comparison of 4sU-Seq log2-transformed fold changes (2 h, 10 Gy IR versus control) for transcripts classified based on either presence or absence of the two top-ranking 7-mers (AGAAGAA, AAGAAGA) in DDX54 binding sites. Median log2-fold changes for both classes are given. (E) Changes in protein synthesis upon IR exposure were measured by pSILAC. Scatter plot of log2-transformed mean H/M ratios for mock and siDDX54 conditions obtained from three replicates of pSILAC experiment. The slope of the linear fit (orange) is shown. Dotted line denotes perfect fit (slope = 1). (F) Box plot of differences in protein synthesis between mock-transfected and DDX54-depleted cells. For proteins with increased synthesis upon IR (H/M ratio >1), the ΔH/M ratios between mock and siDDX54 conditions were computed. The absolute number of proteins encoded by transcripts containing either DDX54-bound or unbound introns, or other proteins with no DDX54 binding information, are shown. A Wilcoxon rank-sum test was used for between-group comparisons.











