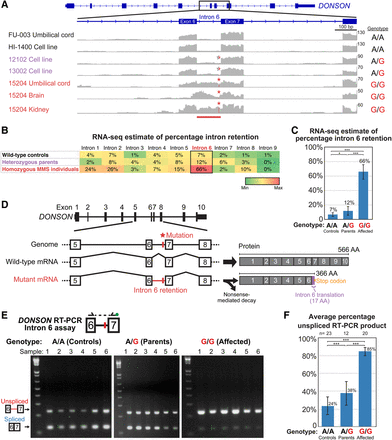
RNA-seq identifies an intron-retention splicing defect associated with an intronic variant in DONSON. (A) RNA-seq read coverage for representative samples shows aberrant retention of intron 6 of DONSON (red bar) in affected individuals associated with the c.1047-9A>G noncoding mutation (Chr 21: g.33582064:T>C; red asterisk). The interval shown is Chr 21: 33,580,994–33,583,594 (GRCh38/hg38), illustrated in the reverse strand direction. The genotype of each sample is shown on the right. Homozygous MMS, heterozygous parent, and wild-type control sample names are colored red, purple, and black, respectively. Read coverage graph Y-axes are scaled (numbers on right side of Y-axis) to show the maximum coverage of each sample in the interval. (B) RNA-seq quantification of intron retention for each intron of DONSON, based upon pooling all the RNA-seq samples of each genotype (see Supplemental Methods for details). Cells are shaded green to red according to the percentile between the minimum and maximum values in the table. The aberrant retention of intron 6 is highlighted. The table also shows the mild increase in intron 6 retention in heterozygous parents relative to wild-type controls and the baseline low-level retention of intron 6 in controls relative to other introns. MMS individuals also showed a trend of increased retention of other introns upstream of intron 6, suggesting that impaired splicing of intron 6 might affect splicing of other introns; however, the mechanism by which this would occur is unclear. (C) RNA-seq quantification of intron 6 retention calculated as in Figure 3B. Error bars are 95% confidence intervals (see Supplemental Methods). All group comparisons were significant: controls versus parents: P = 0.03; controls versus affected: P < 10−15; parents versus affected: P < 10−15 (Fisher's exact test with Holm multiple comparisons adjustment). (D) Schematic of the intron retention splicing defect caused by the c.1047-9A>G (Chr 21: 33582064 T>C) mutation in intron 6 in microcephaly-micromelia syndrome. Retention of intron 6 would lead to either nonsense-mediated decay of the transcript due to the stop codon within intron 6 or to a truncated protein. On the right are the predicted wild-type and truncated mutant proteins and their amino acid (AA) lengths. Translation of the first part of the aberrantly retained intron 6 creates 17 amino acids followed by a premature stop codon. (E) RT-PCR spanning from exon 6 to exon 7 of DONSON (top schematic) in various tissues confirms increased retention of intron 6 in MMS samples, which are homozygous for the Chr 21: 33582064 T>C mutation, compared to heterozygous parents and wild-type controls (unspliced transcript with intron 6: 230 bp; spliced transcript: 121 bp). Shown here are six representative samples for each genotype. (Note that the variant is A>G in the DONSON transcript strand and T>C in the genomic plus strand). See Supplemental Figure 2A for RT-PCR gel images of all assayed samples. The exon 7 PCR primer contains a FAM fluorescent label (green circle) for quantification of PCR products (Fig. 3E). Wild-type (T/T) samples: 1- FU-009 umbilical cord; 2- FU-006 umbilical cord; 3- FU-004 umbilical cord; 4- fetal liver; 5- fetal brain; 6- cerebellum. Heterozygous (T/C) parent samples: 1- 15603 cell line; 2- 15602 cell line; 3- 15202 cell line; 4- 15201 cell line; 5- 15202 blood sample a; 6- 15202 blood sample b. Homozygous (C/C) MMS samples: 1- 15204 brain (RNAlater); 2- 15204 brain (fresh-frozen sample a); 3- 15204 brain (fresh-frozen sample b); 4- 15204 heart (RNAlater); 5- 15204 heart (fresh-frozen); 6- 15204 kidney (RNAlater). (F) Quantification of the RT-PCR intron 6 retention assay products. RT-PCR was performed with the PCR primer for exon 7 containing a FAM-fluorescent label allowing quantification of the RT-PCR products with a capillary electrophoresis DNA analyzer (see Methods). Percentage unspliced RT-PCR product was calculated as [Area of unspliced band]/[Area of unspliced band + Area of spliced band], and averaged across all samples of each genotype (number of samples in each group is shown on top). Groups were significantly different from each other (Controls versus Parents, P = 0.005; Controls vs. Affected, P < 10−22; Parents vs. Affected, P < 10−7; two-tailed unpaired t-test). Importantly, note that this measurement can be used to evaluate relative splicing differences between genotypes but is not an absolute measurement of splicing, since the PCR amplification efficiencies of the unspliced and spliced products differ. See Supplemental Figure 2A for percentage unspliced RT-PCR product of all assayed samples and Supplemental Figure 2B for percentage unspliced RT-PCR product summarized by tissue type for brain, umbilical cord, and cell lines/blood leukocytes.











