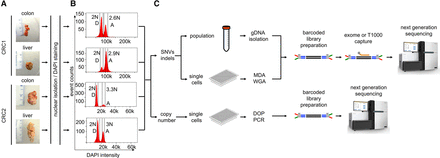
Single-cell and bulk population experimental workflow. (A) The frozen primary tumors and liver metastases from two CRC patients were dissociated into nuclear suspensions and stained with DAPI. (B) Single nuclei and populations of cells were gated and flow-sorted by ploidy distribution. (C) To detect mutations, single nuclei were amplified by multiple-displacement-amplification (MDA), and libraries were captured using the T1000 cancer gene panel, while copy number detection was performed on single nuclei using DOP-PCR. Millions of cells were isolated in parallel for standard exome sequencing. Barcoded libraries were constructed and captured for targeted cancer gene panels or exome panels. Libraries were pooled for next-generation sequencing on the Illumina platform.











