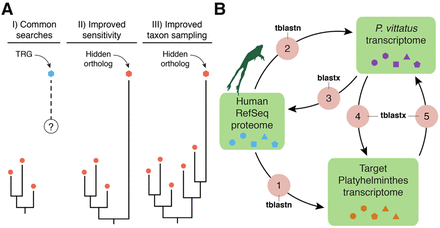
Hidden orthologs and the Leapfrog pipeline. (A) Taxonomically restricted genes (TRGs) are genes with no clear orthology relationship (dashed line and question mark) to other known genes (e.g., orthology group of red dots). Improved sensitivity in the detection methods and/or improved taxon sampling can help uncover hidden orthology relationships, thus referring to these former TRGs as hidden orthologs. (B) The Leapfrog pipeline performs a series of reciprocal BLAST searches between an initial well-annotated data set (e.g., human RefSeq), and a target and a “bridge” transcriptome. First, Leapfrog performs BLAST against the human RefSeq and the target (1) and the “bridge” transcriptome (2) and identifies reciprocal best-hit orthologs between the “bridge” and the human RefSeq proteins (3). These annotated genes of the “bridge” are then used to find orthologs in the target transcriptomes by reciprocal best BLAST hits (4 and 5). If these two pairs of reciprocal best BLAST hit searches are consistent between them, the gene in the target transcriptome is deemed a hidden ortholog. Colored shapes within green boxes represent different sequences of each data set.











