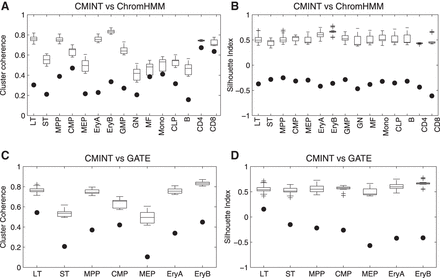
Comparison of CMINT against ChromHMM and GATE on 15 cell types of the hematopoiesis lineage. Cluster coherence (A) and silhouette index (B) of clusters generated by using ChromHMM and CMINT. The filled circles represent the cluster coherence and silhouette index values for ChromHMM. The box plots represent the values obtained using CMINT on 20 different random initializations. Cluster coherence (C) and silhouette index (D) of clusters generated by using GATE and CMINT on one branch of hematopoietic tree. The filled circles represent the silhouette index and cluster coherence values for GATE. The box plots represent the values obtained by CMINT for 20 different runs of the algorithm.











