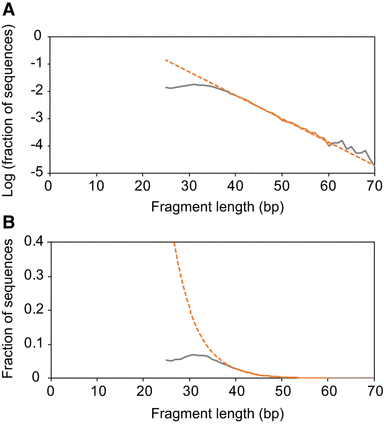
DNA fragment size distribution reconstructed from a hominin femur fragment from Sima de los Huesos. Sequences ≥25 bp from putatively endogenous ancient DNA fragments were isolated from published sequence alignments to the human reference genome (Meyer et al. 2016) by requiring a signal of deamination to be present, i.e., a terminal C to T substitution, and removing alignments with substitutions other than C to T. The first filter depletes human contamination, whereas the second reduces the impact of spurious alignments of microbial sequences. (A) Log-transformed fragment size distribution. Fragment sizes between 40 and 60 bp provide the best fit to an exponential model of DNA decay (dashed line). (B) The same fragment size distribution plotted on a linear scale to visualize more clearly the underrepresentation of short DNA fragments.











