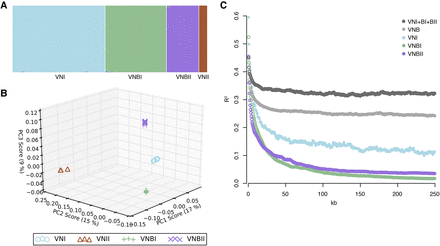
C. neoformans var. grubii is separated into four distinct nonrecombining lineages: VNI, VNII, VNBI, and VNBII. (A) Results of STRUCTURE analysis (k = 4) shows separation of all four lineages. (B) Top three principal components from PCA. The third principal component clearly separates VNBI from VNBII. (C) Decay of linkage disequilibrium (LD) shows similar rates of recombination within groups and lack of recombination between groups. LD (R2) was calculated for all pairs of SNPs separated by 0–250 kb and then averaged for every 500 bp. LD values for each window were then calculated by averaging over all pairwise calculations in the window. The chromosome with the mating type locus was excluded from the calculations.











