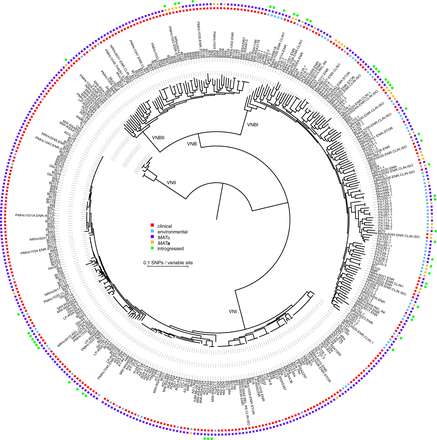
Phylogenomic analysis reveals a deep split within the VNB lineage. We propose that VNB be divided into two lineages: VNBI and VNBII. The phylogeny was estimated from 1,069,080 segregating sites using RAxML (Stamatakis 2014), and the tree was rooted with VNII as the outgroup. All lineages (VNI, VNII, VNBI, and VNBII) had 100% bootstrap support. Isolation source (clinical versus environmental), mating type (MATα and MATa), and presence of ≥50 kb of introgressions from other lineages are also shown. VNBI samples were enriched for environmental isolation sources relative to VNBII (P < 0.0001, Fisher's exact test).











