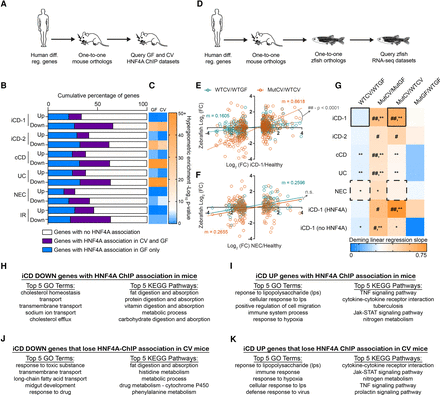
Microbiota suppression of HNF4A activity is highly correlated with genes and intestinal processes suppressed in human IBD and conserved in zebrafish. (A) Flow chart showing the experimental design and filters used to identify IBD, NEC, or IR gene orthologs associated with mouse HNF4A ChIP sites. (B) Bar chart showing the proportion of HNF4A associations in GF and CV mouse jejunal IECs near human-to-mouse one-to-one gene orthologs differentially regulated in human pediatric ileal Crohn's disease (iCD-1), adult iCD (iCD-2), adult colonic Crohn's disease (cCD), adult ulcerative colitis (UC), neonatal necrotizing enterocolitis (NEC), or insulin-resistance (IR). (C) Heat map representing the −log10 (P-value) of the enrichment of GF or CV HNF4A-associated genes that are differentially regulated genes in the indicated IBD data sets. Log10 P-values were calculated using a hypergeometric enrichment analysis and converting all HNF4A ChIP associated mouse genes to human orthologs (GF = 5863 genes and CV = 2119 genes). (D) Flow chart showing the experimental design and filters used to identify correlations between gnotobiotic WT or mutant zebrafish gene expression and gene orthologs differentially expressed in human IBD or NEC. Example of Deming linear regression analysis showing the correlation of log2 (FC) between WTCV/WTGF (E) or MutCV/WTCV (F) zebrafish and pediatric iCD or NEC. m = slope of the line. (G) Heat map representing slopes of Deming linear regression lines showing positive correlative relationships between the log2 gene expression fold changes of one-to-one orthologs from human diseases compared to log2 fold changes in zebrafish WTCV/WTGF, MutCV/MutGF, MutCV/WTCV, and MutGF/WTGF. Because loss of hnf4a function in zebrafish appeared to resemble more closely the iCD signature than cCD or UC, we performed pairwise comparisons of gene orthologs that are (1) differentially regulated in human iCD and (2) have a mouse HNF4A ChIP association (iCD-1 [HNF4A]) or do not have a mouse HNF4A ChIP association (iCD-1 [no HNF4A]). Hash signs indicate slope of Deming linear regression lines is significantly greater than WTCV/WTGF comparison. (#) P < 0.05, (##) P < 0.0001. Asterisks indicate slope of Deming linear regression line is significantly greater than MutGF/WTGF. (*) P < 0.05, (**) P < 0.001. Solid boxes correspond to slope of lines in panel D, and dashed boxes correspond to slope of lines in panel E. (H–K) The top five GO terms and the top five KEGG pathways for indicated gene lists.











