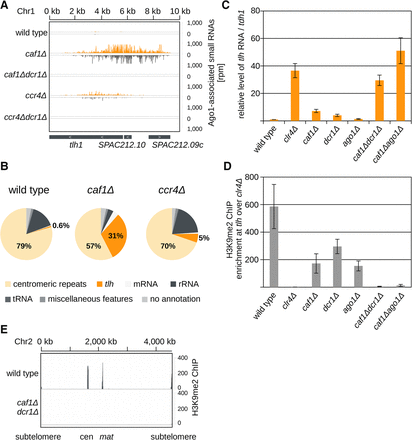
Caf1 and RNAi are required for heterochromatin formation at transcribed regions. (A) Endogenously tagged Argonaute-bound small RNA reads in the indicated cells were plotted over the subtelomeric region. The location of genes is indicated as gray boxes below the small RNA peaks. Reads from + and − strands are depicted in orange and gray, respectively. Scale bars on the right denote small RNA read numbers normalized per one million reads. tlh is present at subtelomeres of both arms of Chromosomes 1 and 2 (Mandell et al. 2005; Hansen et al. 2006). (B) Classification of Argonaute-bound small RNAs from wild-type, caf1Δ, and ccr4Δ cells. Pie charts illustrate percentages for the individual small RNA classes relative to the total number of reads for each strain. Argonaute-bound subtelomeric siRNAs are increased by more than 50-fold in caf1Δ cells compared with the wild type. (C) Quantification of subtelomeric tlh transcripts in indicated strains by RT-qPCR. In caf1Δdcr1Δ and caf1Δago1Δ cells, silencing of subtelomeric repeats is lost. Error bars, SE of more than seven independent experiments. For caf1Δdcr1Δ and caf1Δago1Δ, experiments of two independent colonies were averaged, respectively. Reverse transcription was performed with specific primers; the wild type was set to one. (D) ChIP experiment showing that H3K9me2 is lost at subtelomeric tlh repeats in caf1Δdcr1Δ cells. Error bars, SE of at least four independent experiments. For caf1Δdcr1Δ and caf1Δago1Δ, data of two independent colonies were averaged, respectively. clr4Δ was set to one. (E) ChIP-seq showing that H3K9me2 is lost at all constitutive heterochromatic loci in caf1Δdcr1Δ cells. Scale bars on the right denote read numbers per million reads normalized to the TAS region (Chr 2: 4534 kb–4538 kb) where H3K9me2 is not lost in the mutant.











