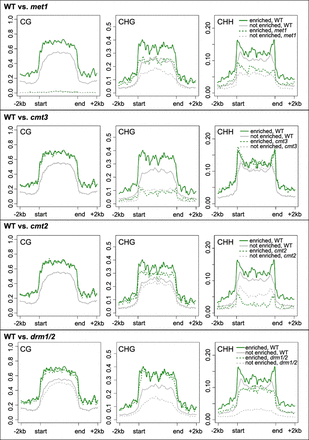
Figure 6.
Comparison of DNA methylation over TEs. Patterns of TE DNA methylation (CpG, CHG, CHH) in wild-type (WT) and mutants. The grouping of TEs is according to the enrichment results of NUP1:GFP RE-ChIP-seq from 30-d-old leaf tissues. The methylation ratio is calculated in 100-bp windows. The signal over each TE is linearly transformed so that the boundaries of all TEs are aligned.











