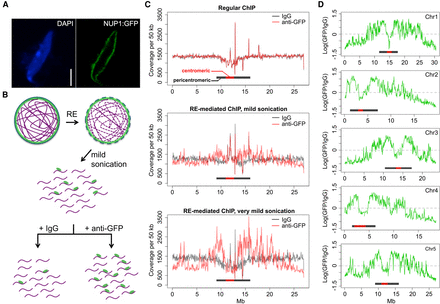
Identification of chromatin located at the nuclear periphery by RE-ChIP. (A) Localization of the NUP1:GFP protein in an Arabidopsis nucleus: (scale bar) 2 µm. (B) Procedures for RE-mediated ChIP with NUP1:GFP (green). Chromatin (purple lines) fragmentation and isolation are conducted with a combination of RE (restriction enzyme) digestion and mild sonication. (C) Normalized sequence coverage (50-kb window size) on Chromosome 5 from various ChIP experiments. The horizontal bars depict pericentromeric regions, within which centromeric regions are highlighted in red. (D) NUP1:GFP RE-mediated ChIP-seq signal (50-kb window size), represented as the log2 value of the ratio between normalized anti-GFP and IgG coverage, over all five chromosomes. Horizontal bars indicate the centromeric/pericentromeric regions, as in C.











