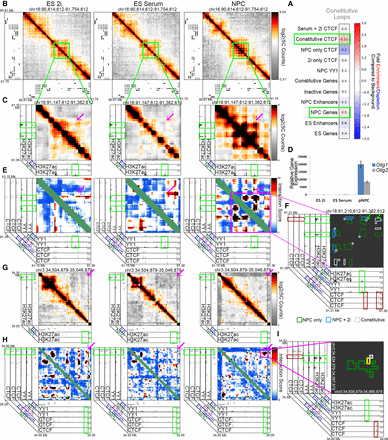
YY1 connects neural regulatory elements nested within and adjacent to a framework of constitutive CTCF-mediated interactions. (A) Fold enrichment/depletion of chromatin regulatory elements in the constitutive looping class compared to background interactions. P-values are computed with Fisher's exact test and listed in each entry. (B,C) Relative interaction frequency heatmaps of ∼1 Mb region (B) and ∼200 kb region (C) surrounding the Olig1 and Olig2 genes in ES 2i, ES serum, and NPCs. Heatmaps in C are overlaid with ChIP-seq tracks of H3K27ac in ES serum cells and NPCs. (D) Relative gene expression of Olig1 and Olig2 genes across the ES 2i, ES serum, and NPC cellular states. (E) Zoom-in interaction score heatmaps of looping interactions between the Olig1 and Olig2 genes and surrounding putative NPC enhancers (green boxes). (F) Zoom-in cluster map of classified looping interactions at Olig2 and Olig1 with NPC only (green), serum+NPC (blue), and constitutive class looping interactions (gray). (G–I) Heatmaps and cluster map at different length scales around the Sox2 gene in ES 2i, ES serum, and NPCs. Zoom-in heatmaps of relative interaction frequencies (G) and background corrected interaction scores (H) across ∼500 kb downstream from Sox2. Relative interaction frequency heatmaps are overlaid H3K27ac tracks. Interaction score heatmaps are overlaid with ChIP-seq tracks of YY1 and CTCF across cell types. The Sox2 gene is colored green. (I) Zoom-in classified cluster map of a ∼100-kb window around a Sox2-enhancer interaction with NPC only (green), serum+NPC (yellow), and constitutive classified looping interactions (gray), overlaid on ChIP-seq tracks.











