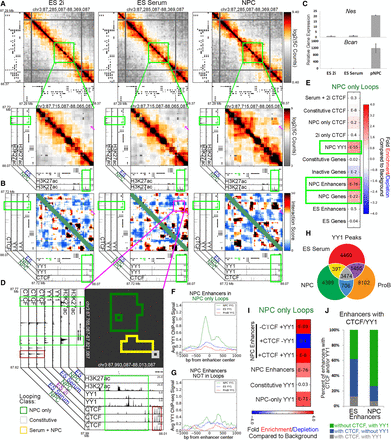
YY1 is enriched at NPC-specific enhancers that form developmentally regulated loops. (A) Relative interaction frequency heatmaps of the global view of 1 Mb surrounding Nes (top), and zoom in of 400 kb surrounding nestin with putative NPC enhancer annotations (bottom). Nes (upstream) and Bcan (downstream) genes are colored green. (B) Zoom-in interaction score heatmaps of the Nes/Bcan genes interacting with a downstream putative NPC enhancer. Heatmaps are overlaid with ChIP-seq tracks of CTCF in NPCs and YY1 in ES serum and NPCs. The Nes (upstream) and Bcan (downstream) genes are colored green. (C) Relative gene expression of Nes and Bcan across ES 2i, ES serum, and NPC cellular states. (D) Interaction cluster outlines of the loop boxed in magenta in B. Plot is overlaid with ChIP-seq tracks of H3K27ac, YY1, and CTCF in the ES 2i, ES serum, and NPC conditions. Cluster outline classifications include NPC-only (green), serum+NPC (yellow), and constitutive (gray). (E) Fold enrichment/depletion of the presence of chromatin features in NPC-only interaction class compared to the presence in background. P-values are computed with Fisher's exact test and listed in each entry. (F,G) YY1 ChIP-seq signal in NPCs (green), ES serum (blue), and ProB cells (red), centered at (F) putative NPC enhancers at the base of NPC-only loops, and (G) NPC enhancers that do not fall at the base of any looping interactions. (H) YY1 binding sites parsed by their occupancy across ES cells, NPCs, and ProB cells. (I) Fold enrichment/depletion of YY1 peak classes and NPC enhancers parsed based on the presence/absence of CTCF/YY1 in NPC-only loops compared to their presence in background interactions. (J) Stacked bar plot of the breakdown of ES and NPC enhancers that are bound with confidence by a combination of CTCF and/or YY1.











