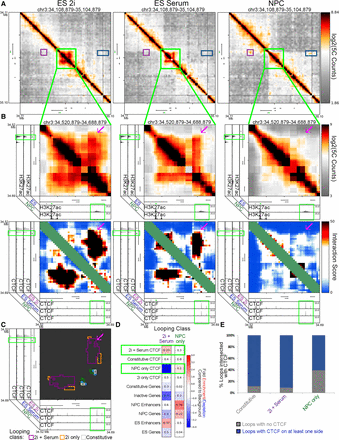
Pluripotency interactions that disengage in multipotent NPCs display reduced CTCF occupancy. (A) Global view of relative interaction frequency heatmaps of 1 Mb surrounding the Sox2 gene. (B) Zoom in highlighting a strong pluripotency-specific looping interaction between Sox2 and an ES-specific enhancer. CTCF binds at both loop anchors (note green boxes). Heatmaps include relative interaction frequency (top) and background corrected interaction score (bottom). The Sox2 gene is colored green. (C) Classified interaction clusters are plotted above relevant ChIP-seq tracks. (D) Fold enrichment/depletion of chromatin features in 2i+serum and NPC-only looping interaction classes compared to presence in background interactions. P-values included in each entry are calculated using Fisher's exact test. (E) Stacked bar plot contrasting the proportion of loops connected by CTCF in one or both anchoring fragments versus not anchored by CTCF.











