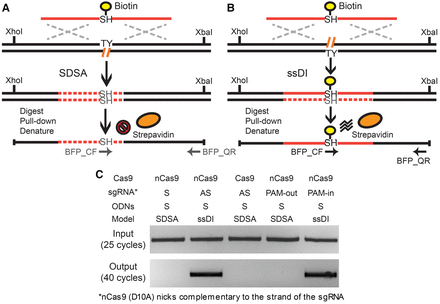
The biotin pull-down assay. (A,B) Schematic illustration of the biotin pull-down assay via the SDSA (A) and ssDI (B) pathways. In the case of ssDI (B), the biotinylated ODN is predicted to be incorporated into the target genomic locus, whereas in SDSA (A), it should not. The XhoI- and XbaI-digested genomic fragments covalently linked to biotin can be enriched using streptavidin beads under denaturing conditions. The primers BFP_QF and BFP_QR can specifically amplify these genomic fragments with ODN incorporation but not free ODN donors. Biotin, yellow circles; streptavidin, orange ovals; genomic DNA, black lines; ODN sequence, solid red lines; DNA synthesis, dashed red lines; chromophore sequence, TY and SH; genomic lesions, hatched orange lines; homology regions, dashed silver crosses; restriction sites, XhoI and XbaI; PCR primers, horizontal arrows. (C) Results of the biotin pull-down assay. After transfecting with biotinylated ODNs (BFP_S90_Biotin) (Supplemental Fig. S8), the BFP-positive cells were enriched using FACS sorting. The genomic DNA of the BFP-positive cells was digested with XhoI and XbaI. The fragments covalently linked to biotin were pulled down with streptavidin beads in denaturing conditions, PCR-amplified using one internal primer and one flanking primer of the ODN donors (BFP_QF and BFP_QR) (Supplemental Fig. S8), and detected on an agarose gel.











