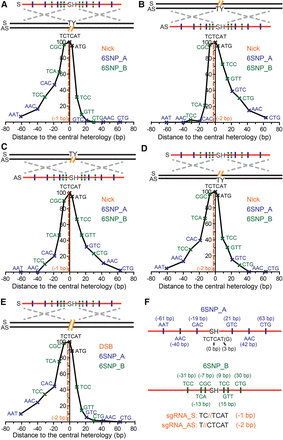
Conversion tracts of single-nick– and DSB-induced HDR using ODN donors. (A–E) The conversion tracts of single-nick–induced HDR using ODN donors complementary to (A,B) or the same strand of (C,D) the nick, and DSB-induced HDR (E). The conversion tracts were compiled by overlaying the retention frequency of each SNP in both 6SNP_A and 6SNP_B donors. The positions of the SNPs and predicted genomic lesions are labeled in reference to the center of the chromophore sequence. Schematic elements are colored as follows: genomic DNA, black; genomic lesions, orange; ODNs, red; chromophore sequences, TY and SH; SNPs on the 6SNP_A ODNs, blue; SNPs on the 6SNP_B ODNs, green; homology regions, dashed silver crosses; strandedness of DNA, S and AS. (F) Schematics of the ODN donors with synonymous SNPs. The SNPs are represented as the central nucleotide of the trinucleotides in the S orientation, and the distance between the SNPs and the center of the SH sequence (TCTCAT) is labeled. Because the last T in the SH sequence is a wobble nucleotide, it could be used as a SNP as well and is represented as the ATG at position +3. ODNs with six distributed or clustered SNPs are labeled as 6SNP_A (top) and 6SNP_B (bottom), respectively. The sequences shown at the bottom of the panel emphasize that the S nick is made at position −1 and the AS nick at position −2. The actual sequences of the ODNs can be found in Supplemental Figure S8.











