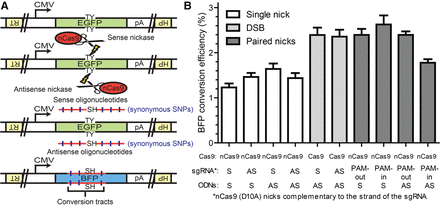
The EGFP to BFP conversion system in the HPRT1-EGFP antisense cell line. (A) Schematics of the EGFP to BFP conversion system in the HPRT1-EGFP antisense cell line. An EGFP expression cassette was knocked into the HPRT1 locus of the HCT116 cell line (only the antisense orientation is diagrammed here for the sake of ease of presentation). Genomic lesions could then be introduced near the chromophore (TY residues) of the EGFP sequence using CRISPR/Cas9 cleavase, nickases, or dual nickases. HDR repair of the genomic lesions using ODN donors containing the sequence of the SH residues leads to the conversion of EGFP to BFP. The ODN donors also contained synonymous SNPs that could be coconverted with the sequence of the SH residues. Schematic elements: EGFP, green boxes; BFP, blue boxes; HPRT1, inverted yellow boxes; sequences of the chromophore residues, TY and SH; the cytomegalovirus promoter, CMV; the polyadenylation sequence, pA; Cas9 variants, red ovals with scissors; genomic lesions, lightning bolts; ODN donors, horizontal red lines; synonymous SNPs, vertical blue hash marks. (B) The efficiency of BFP conversion in the HPRT1-EGFP antisense cell line. The wild-type Cas9 and D10A variant are labeled as Cas9 and nCas9, respectively. The strandedness (S, sense; AS, antisense) of the sgRNA and ODNs (for this experiment without synonymous SNPs) are labeled with respect to the coding sequence of EGFP. Note that the Cas9 D10A variant nicks the strand complementary to that of the sgRNA. All data are shown as the mean ± SEM of three biological replicates.











