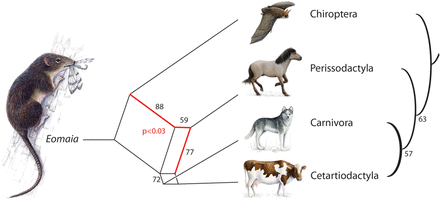
Phylogenetic network and phylogenetic tree reconstruction based on retrotransposon markers for the four investigated laurasiatherian orders. Neighbor-net (SplitsTree) analysis (left) and the most parsimonious tree reconstruction (Dollop in Phylip) (right) were conducted based on the presence/absence patterns of 102 retrotransposon insertions. Black numbers represent bootstrap values. The red number indicates the χ2 significance value from the four-lineage insertion likelihood test (Supplemental Material S1) favoring the hybridization scenario merging horse with the bat and dog or dog/cow ancestor (red lines in the SplitsTree). An imaginary reconstruction of the Eomaia, an assumed ancestor of placentals, based on a description of Ji et al. (2002) is presented at the root of the “tree.”











