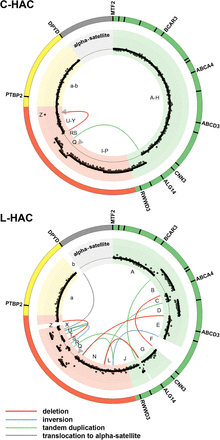
Structural variation of C-HAC and L-HAC. Circos plots show four HAC segments and the structural rearrangements between them. The gene-rich segment is indicated in green; gene-poor, gene desert, and alpha satellite repeat centromeric segments are depicted in yellow, red, and light gray, respectively. The lost sequences are indicated by white gaps. In the C-HAC plot, the NEO-loxP-3′HPRT1-construct insertion sites are indicated by gray arrowheads; and in the L-HAC plot, the gray arrowheads point to the location of telomeres. The size of the alpha satellite region is not to scale. Dots in the inner colored circle show the logR values per 10-kb bin, solid black lines indicate segments with estimated integer DNA copy number 1, 2, or 3. Chromosome rearrangement signatures are indicated by curved colored lines: deletions, red; tandem duplications, green; inversions, blue; translocations to the centromere region, gray. The letters (A–Z, a-b) indicate names of the blocks specified in Supplemental Table S2. Names of some genes are indicated on the outer colored circle.











