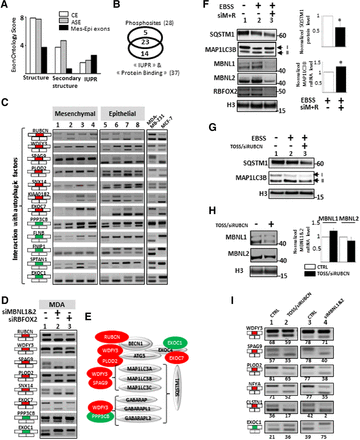
(A) Exons differentially spliced between epithelial- and mesenchymal-like cells (Mes-Epi exons) code for protein segments poorly associated with “structure” and “secondary structure” terms but that do contain unstructured regions (IUPR). (B) 28 and 37 exons out of the 81 selected exons code for protein segments containing experimentally validated phosphosites, IUPRs, and/or “Protein Binding” motifs, respectively; 23 of them contain both phosphosites and protein interacting motifs. (C) RT-PCR performed with total RNAs obtained from four normal epithelial (1 = HEPic, 2 = HPAEPic, 3 = HMEC, 4 = AG01134) and four normal mesenchymal (5 = HMF, 6 = HCFaa, 7 = AG0449, 8 = AG0450) cell lines and from MCF-7 and MDA-MB-231 breast cancer cell lines. The selected genes correspond to genes bearing alternative exons coding for protein interacting with autophagic factors. Red and green rectangles correspond to alternative exons with higher and lower inclusion rate, respectively, in mesenchymal-like cells compared to epithelial-like cells. (D) RT-PCR corresponding to genes with alternative exons and interacting with proteins involved in autophagy, using total RNAs obtained from mesenchymal-like MDA-MB-231 breast cancer cells transfected with control siRNAs (lane 1), siRNAs targeting MBNL1 and MBNL2 (lane 2), or RBFOX2 (lane 3). Red and green rectangles correspond to alternative exons with higher and lower inclusion rate, respectively, in mesenchymal-like cells compared to epithelial-like cells. (E) Genes with exons regulated by mesenchymal cell-enriched splicing factors produce proteins interacting with proteins involved in autophagy. Red and green proteins correspond to genes with alternative exons with higher and lower inclusion rate, respectively, in mesenchymal-like cells compared to epithelial-like cells. (F) Western blot analyses of SQSTM1, MAP1LC3B, MBNL1, MBNL2, and RBFOX2 in control (Earle's Balanced Salt Solution−[EBSS−]) or serum starved (EBSS +) MDA-MB-231 cells transfected with control siRNAs or siRNAs targeting MBNL1, MBNL2, and RBFOX2 (siM+R). H3 (histone H3) is used as a loading control. The quantification of the SQSTM1 Western blot signal and MAP1LC3B mRNA level by RT-qPCR is shown on the right. (*) P-value < 0.05. (G) Western blot analyses of SQSTM1 and MAP1LC3B in control (EBSS−) or serum starved (EBSS +) MDA-MB-231 cells transfected with TOSS and siRNA targeting RUBCN exon 14 (TOSS/siRUBCN). H3 (histone H3) is used as a loading control. (H) MDA-MB-231 cells were transfected with TOSS and siRNA targeting RUBCN exon 14 (TOSS/siRUBCN). Western blot analysis of MBNL1 and MBNL2, with H3 (histone H3) used as a loading control (left panel). RT-qPCR analysis of the MBNL1 and MBNL2 mRNA levels in the same experimental conditions (right panel). (I) RT-PCR analysis using total RNAs extracted from mesenchymal-like MDA-MB-231 cells transfected as described in panel H or transfected with siRNAs targeting MBNL1 and MBNL2 (siMBNL1&2).











