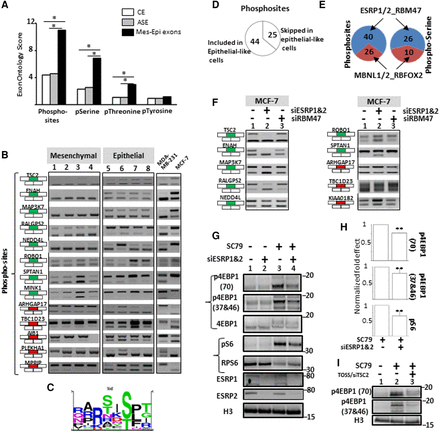
(A) Exons differentially spliced between epithelial- and mesenchymal-like cells (Mes-Epi exons) encode for protein segments that are enriched, when compared to constitutive (CE) or alternative (ASE) exons, for “Phosphorylated residue” (Phosphosites), “O-phospho-L-serine” (pSerine), “O-phospho-L-threonine” (pThreonine) terms but not for the “O4′-phospho-L-tyrosine” (pTyrosine) term. (*) FDR adjusted P value < 0.05. (B) RT-PCR performed with total RNAs obtained from four normal epithelial (1 = HEPic, 2 = HPAEPic, 3 = HMEC, 4 = AG01134) and four normal mesenchymal (5 = HMF, 6 = HCFaa, 7 = AG0449, 8 = AG0450) cell lines and from MCF-7 and MDA-MB-231 breast cancer cell lines. The selected genes correspond to genes bearing alternative exons coding for protein segments containing experimentally validated phosphosites. Red and green rectangles correspond to alternative exons with higher and lower inclusion rate, respectively, in mesenchymal-like cells compared to epithelial-like cells. (C) Sequence logo generated from the “PhosphoSite” website using sequences surrounding experimentally validated phosphorylated residues coded by exons differentially spliced between epithelial- and mesenchymal-like cells. (D) 44 and 25 experimentally-validated phosphorylated residues are encoded within exons more often included and less often included, respectively, in epithelial-like cells. (E) 40 and 26 experimentally validated phosphorylated residues are encoded within exons regulated by epithelial cell-enriched splicing factors (ESRP1, ESRP2, and RBM47) and by mesenchymal cell-enriched splicing factors (MBNL1, MBNL2, and RBFOX2), respectively (left panel). 26 and 10 experimentally validated phosphorylated serine residues are encoded within exons regulated by epithelial cell-enriched splicing factors (ESRP1, ESRP2, and RBM47) and by mesenchymal cell-enriched splicing factors (MBNL1, MBNL2, and RBFOX2), respectively (right panel). (F) RT-PCR performed with total RNAs extracted from epithelial-like MCF-7 cells transfected with control siRNAs (1), siRNAs targeting ESRP1 and ESRP2 (2) or RBM47 (3). The selected genes correspond to genes bearing alternative exons encoding for experimentally validated phosphorylated residues. Red and green rectangles correspond to alternative exons with higher and lower inclusion rate, respectively, in mesenchymal-like cells compared to epithelial-like cells. (G) Western blot analyses of the phosphorylation pattern of proteins involved downstream of the AKT signaling pathway in epithelial-like MCF-7 cells transfected with control siRNAs or siRNAs targeting ESRP1 and ESRP2 and treated, or not, for 1 h with SC79 (AKT kinase activator). The p4EBP1(70) and p4EBP1(37&46) antibodies recognize phosphorylated residues on position 70, 37 and/or 46 of the E4BP1 protein. The pS6 antibody recognizes phosphorylated RPS6 protein. H3 (histone H3) is used as a loading control. (H) Quantification of Western blots shown in panel G. p4EBP1(70) and p4EBP1(37&46) signals were normalized by the signal obtained with an antibody recognizing both phosphorylated and unphosphorylated 4EBP1 protein (4EBP1). Likewise, the pS6 signal was normalized to total RPS6 signal (S6). (**) P-value < 0.005. (I) Western blot analyses of the phosphorylation pattern of 4EBP1 protein in MCF-7 cells transfected, or not, with TOSS and siRNAs targeting TSC2 exon 27 (TOSS/siTSC2) and treated, or not, for 1 h with SC79. H3 (histone H3) is used as a loading control.











