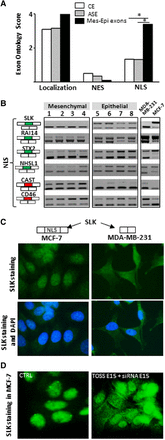
(A) Exons differentially spliced between epithelial-like and mesenchymal-like cells (Mes-Epi exons) code for protein segments that are enriched in “NLS” term when compared to constitutive (CE) or alternative (ASE) exons. (*) P-value < 0.05. (B) RT-PCR performed with total RNAs obtained from four normal epithelial (1 = HEPic, 2 = HPAEPic, 3 = HMEC, 4 = AG01134) and four normal mesenchymal (5 = HMF, 6 = HCFaa, 7 = AG0449, 8 = AG0450) cell lines and from MCF-7 and MDA-MB-231 breast cancer cell lines. The selected genes correspond to genes bearing alternative exons coding for protein segments containing nuclear localization signal (NLS). Red and green rectangles correspond to alternative exons with higher and lower inclusion rate, respectively, in mesenchymal-like cells compared to epithelial-like cells. (C) SLK exon 15 that encodes for a NLS is more often included in MCF-7 than in MDA-MB-231 cells (see panel B). Immunofluorescence of SLK protein indicates that SLK is more restricted to the nucleus in MCF-7 (epithelial-like) than in MDA-MB-231 (mesenchymal-like) cells. (D) Depletion of SLK transcripts that contain exon 15 (TOSS E15 + siRNA E15) leads to a more diffuse SLK staining within transfected MCF-7 cells compared to control (CTRL) cells.











