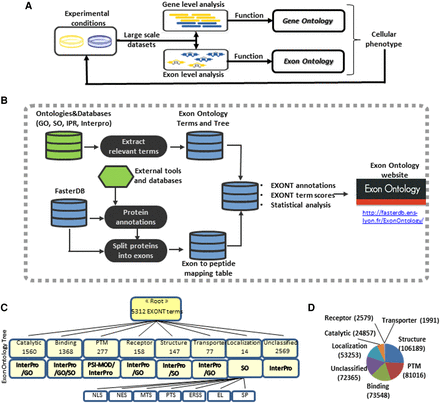
(A) Genome-wide transcriptomic analyses allow identification of genes whose expression levels are modified when comparing two experimental conditions. Looking for the enrichment of Gene Ontology (GO) terms associated with these genes allows prediction of the biological processes and cellular activities that are likely to be impacted by gene expression level modifications. Exon Ontology aims at identifying protein features associated with changes of exon content owing to alternative splicing regulation, which may contribute to cellular phenotypes. Both GO and Exon Ontology predictions can next be addressed by dedicated experimental approaches. (B) The Exon Ontology workflow is based on ontological terms (EXONT terms) that were derived from existing ontologies and databases (e.g., GO, Sequence Ontology, PSI-MOD, and InterPro). Protein features were derived from reference tools and databases and were mapped to annotated genomic exons in the ‘Faster DB’ database. Genomic exons can thus be associated with one or several EXONT terms. A computational suite (Exon Ontology) then calculates a dedicated EXONT term score and looks for potential EXONT term enrichment by statistical analysis. (C) Protein features and domains have been assigned Exon Ontology (EXONT) terms based on existing ontologies and databases as described in panel B. The EXONT terms were organized in an Exon Ontology tree based on height classes of protein features (e.g., catalytic, binding). Each class was divided in categories and contains a more or less large number of associated terms. For example, the “Localization” class was divided into “Nuclear Localization Signal” (NLS), “Nuclear Export Signal” (NES), “Mitochondrial Targeting Signal” (MTS), “Peroxisomal Targeting Signal” (PTS), “Endoplasmic Reticulum Signal Sequence” (ERSS), “Endosomal Localization Signal” (EL), and “Signal Peptide” (SP) categories based on the “Sequence Ontology” resource. (D) Pie chart showing the distribution of functional annotations of human coding exons; more than 170,000 human coding exons are associated with at least one Exon Ontology term. The numbers represent the number of exons associated with each of the main classes of the Exon Ontology terms.











