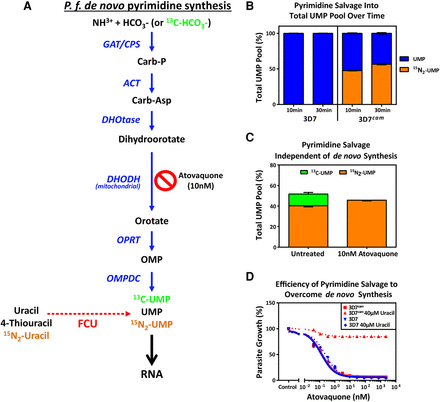
Efficiency of pyrimidine salvage in FCU-GFP expressing P. falciparum. (A) In Plasmodium parasites, pyrimidines are metabolized de novo from bicarbonate and ammonia through a series of enzymatic reactions (blue arrows) resulting in the generation of UMP, which can be incorporated into nascent RNA (black arrow). FCU-GFP allows for salvage of the pyrimidine precursor uracil (red dashed arrow) to UMP, which can then be incorporated into RNA. Inhibition of de novo pyrimidine synthesis is achieved via treatment with the mitochondrial inhibitor atovaquone. (B) 4-TU is efficiently taken up and incorporated into the pyrimidine pool by P.f. 3D7cam parasites grown in the presence of 40 µM 15N2-uracil for 10 and 30 min. Cellular metabolites were detected via LC-MS, and the proportion of total UMP pool that is unlabeled (blue) and labeled with 15N2 (orange) was calculated (n = 2 ± SD performed in triplicate). (C) Flux through de novo pyrimidine synthesis and salvage was measured in treated and untreated parasites by LC-MS after the addition of 13C-bicarbonate (13C-HCO3-) and 15N2-uracil to the culture medium for 30 min. The percentage of UMP derived from de novo synthesis (13C-UMP) and salvage (15N2-UMP) was determined in the presence or absence of the inhibitor atovaquone (10 nM) (n = 2 ± SD performed in triplicate). (D) Early ring stage P. falciparum 3D7 and 3D7cam were exposed to titrating concentrations of atovaquone with or without supplementation of 40 µM uracil to the medium. Parasite survival was determined using a traditional 48-h SYBR-green growth inhibition assay and plotted as an average of three technical replicates (±SD).











